Anders Sejr Hansen, PhD

Dynamics of 3D Genome Structure and Function
The Hansen lab develops new methods to resolve 3D genome structure at high spatiotemporal resolution to understand how DNA looping and 3D folding regulates gene expression in health and disease.
Staff
Research:
Exquisite regulation of gene expression in time and space underlies essentially all biological processes. However, understanding gene regulation in mammals is challenging because mammalian genomes are enormous and largely non-coding. Specifically, mammalian genes are controlled by enhancers that can be hundreds of kilobases away from the genes they control. How the cell ensures that the right enhancer loops to and activates the right gene in 3D is not well understood. We are broadly interested in understanding how the 3D organization of the genome is regulated and how this 3D organization in turn regulates gene expression.
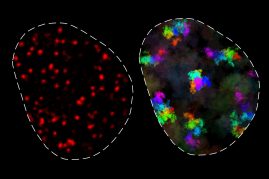
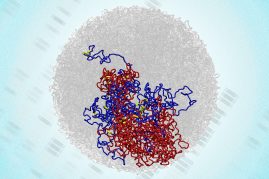
To understand the relationship between 3D genome structure and function, we develop new technologies. We develop new super-resolution and single-molecule imaging methods that allow us to track single proteins as well as DNA looping and nascent transcription in real-time inside of living cells at high spatiotemporal resolution. Moreover, we develop new 3D genomics methods to map 3D genome structure at ultra-high resolution. We also develop new computational methods drawing on diverse fields including image processing and computer vision, machine learning, Bayesian inference, mathematical modeling, and polymer modeling. Finally, we integrate these approaches with traditional methods from genomics, biochemistry, genome-editing, stem cell differentiation and beyond. Current areas of interest include:
- Understanding the molecular and biophysical mechanisms that regulate 3D genome organization
- Understanding how 3D folding and DNA looping controls gene expression
- Understanding how dysregulation of 3D genome structure causes disease
- Developing new microscopy methods for tracking single molecules inside of living cells
- Developing new genomics methods for mapping 3D genome structure and function
- Developing computational models of 3D genome organization
- Bottom-up synthetic biology approaches to test our understanding and to control endogenous regulation
Biography:
Anders obtained his undergraduate and Master’s degree in Chemistry at Oxford University in 2010. He received his PhD in Chemistry and Chemical Biology from Harvard University in 2015, where he worked with Erin O’Shea and applied systems biology approaches to understand how cells can encode and transmit information in the dynamics of transcription factor activation. For his post-doc at UC Berkeley with Robert Tjian and Xavier Darzacq, Anders developed new imaging approaches for dissecting the dynamics of architectural proteins with single-molecule resolution in living cells. Anders joined MIT as an Assistant Professor of Biological Engineering in early 2020, where his lab is focused on understanding and engineering 3D genome structure and function. Anders has won several awards including an NIH K99 Pathway to Independence award (2019), NIH Director’s New Innovator award (2020), a Pew-Stewart Scholar for Cancer Research award (2021), an NSF CAREER award (2024), and an NIH Director’s Transformative Research Award (2024).